

instituto de biologia molecular e celular | institute for molecular and cell biology
Research Results
Lessons on the evolutionary forces and molecular mechanisms that drive the divergence of populations and species, and ultimately speciation, can be taken from the analyses of genes that contribute to the cessation of gene flow between populations. Barriers to gene flow can arise at multiple prezygotic and postzygotic life-history stages. In flowering plants, one of the most common post-pollination prezygotic barriers, is gametophytic self-incompatibility (GSI). In this widespread system, when the S-pollen specificity matches that of the S-pistil, the pollen is recognized as “self” and is rejected by the pistil. This system is ideal to address general principals on how life can innovate, since it has reappeared independently several times during plant evolution.
Although in eudicots there is evidence for a single S-RNase based GSI origin, in Rosaceae species, at the S-RNase gene, different amino acid positions are involved in specificity determination. Moreover, levels of recombination, the rate at which new specificities arise, the number of ancestral lineages and the degree of specificity sharing between closely related Rosaceae species are also different. Furthermore, in Prunus the S-pollen gene is a single gene. In contrast, in Pyrinae (former Maloideae) multiple genes determine S-pollen specificity. Different mechanisms may thus, be used to achieve the rejection of incompatible pollen in different plant families.
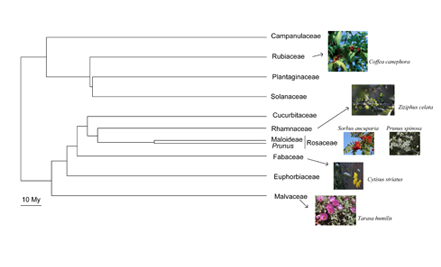
IMAGE: Plant families presenting S-RNase based GSI. Species under study are shown.
Future Research
To highlight the features of the ancestral GSI system we are characterizing the S-RNase and S-duplicated genes in Fabaceae, Rhamnaceae, Malvaceae, and Rubiaceae species. The current hypothesis is that the common ancestor of these plant families possessed an S-RNase-based GSI system. Nevertheless, remarkable differences are observed in Rosaceae species such S-RNase gene intron number, number of S-pollen genes, or competitive interaction (the breakdown of SI in heteroallelic pollen) documented in Malus but, that does not exist in Prunus. The alternative hypothesis, that different gene family members have being recruited for GSI in these familes is thus, being tested.
In Prunus, the S-pollen protein is assumed to protect self S-RNases from being inhibited by a general S-RNase inhibitor. We are addressing the predictions of the general inhibition model such as: testing the interaction of the two proteins, expression and purification of recombinant Prunus SFB and S-RNase, determining the three-dimensional structure of the S-RNase, SFB and their complex by X-ray crystallography, and identification of SFB and S-RNase binding partners.
Selected references:
Vieira J, Morales-Hojas R, Santos RA, Vieira CP. 2007. Different positively selected sites at the gametophytic self-incompatibility pistil S-RNase gene in the Solanaceae and Rosaceae (Prunus, Pyrus, and Malus). J Mol Evol. 65:175-85
Vieira J, Fonseca NA, Vieira CP. 2008a. An S-RNase-based gametophytic self-incompatibility system evolved only once in eudicots. J Mol Evol. 67:179-90.
Vieira J, Teles E, Santos RAM, Vieira CP. 2008b. Recombination at Prunus S-locus region SLFL1 gene. Genetics 180:483-91.
Tsukamoto T, Potter D, Tao R, Vieira CP, Vieira J, Iezzoni AF. 2008. Genetic and molecular characterization of three novel S-haplotypes in sour cherry (Prunus cerasus L.). J Exp Bot. 59:3169-85.
Vieira J, Fonseca NA, Vieira CP. 2009. RNase based gametophytic self-incompatibility evolution: questioning the hypothesis of multiple independent recruitments of the S-pollen gene. J Mol Biol. 69:32-41.
Vieira J, Ferreira PG, Aguiar B, Fonseca NA, Vieira CP. 2010. Evolutionary patterns at the RNase based gametophytic self - incompatibility system in two divergent Rosaceae groups (Maloideae and Prunus). BMC Evol. Biol. 10:200.

Rocha, Sara
sara.rocha@i3s.up.pt
Bastos, Bárbara
barbara.bastos@ibmc.up.pt
Dias Sousa, André
andre.sousa@i3s.up.pt
Home | Site Map | Contacts | Credits | Privacy & Cookies | WHISTLEBLOWER CHANNEL | Intranet | Social Networks |
